GENEIOUS PRIME
CRISPR gRNA Design Software
Powerful software designed for CRISPR makes it quick and easy to find sites, design guide RNAs and analyze your editing results
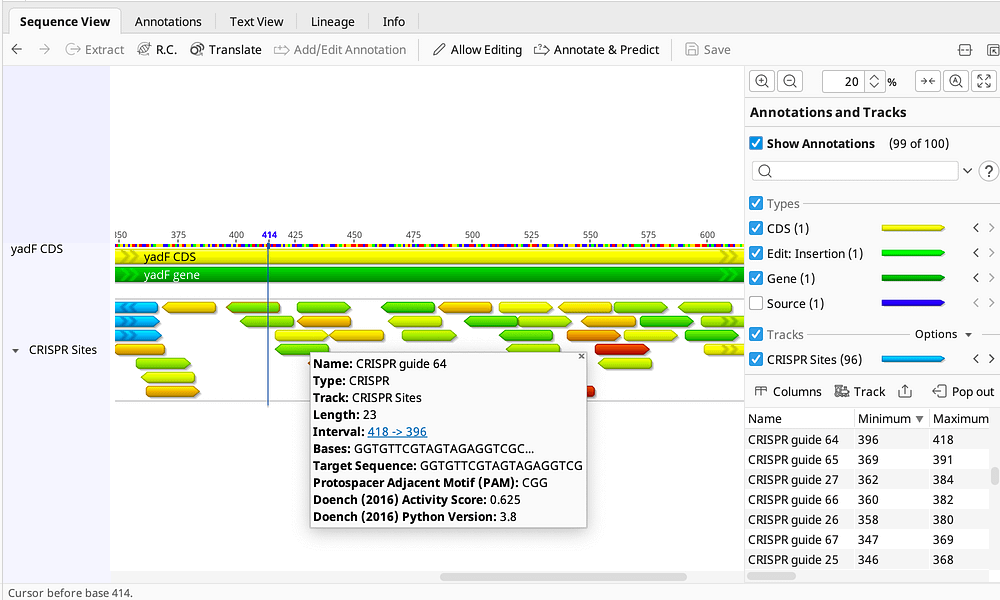
Find CRISPR Sites
- Locate CRISPR target sites and score them within a selected region of a genome or a stand alone gene sequence
- Batch discovery and scoring of CRISPR sites on sequence lists or groups of sequence documents
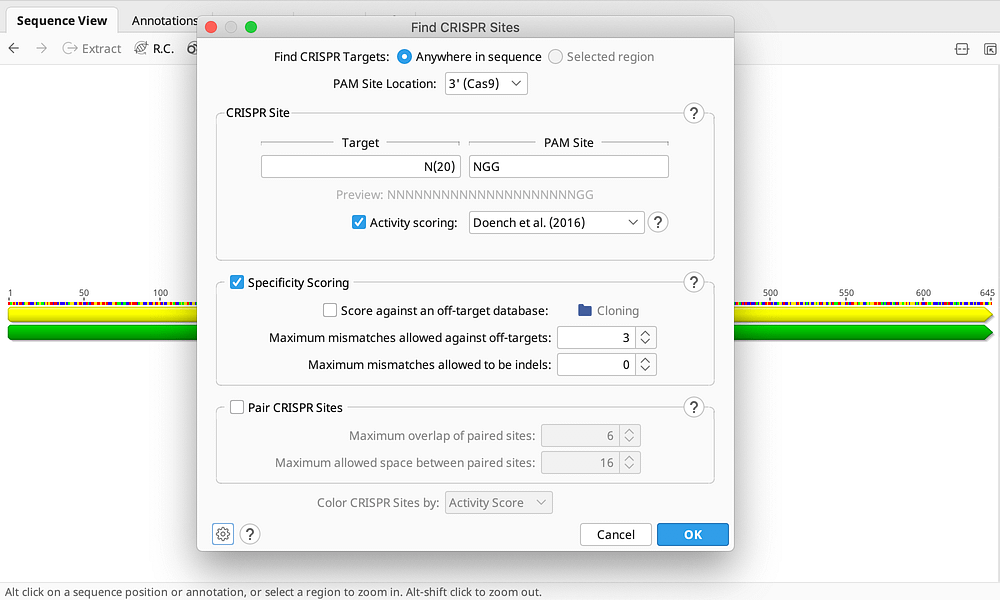
- Support for 3’ (Cas9) and 5’ (Cpf1) PAMs with customizable guide length and motif
- Automatically identify optimal pairs of CRISPR sites
Evaluate Candidates
- Assess sites quickly with innovative heatmap-style annotations based on your preferred scoring method
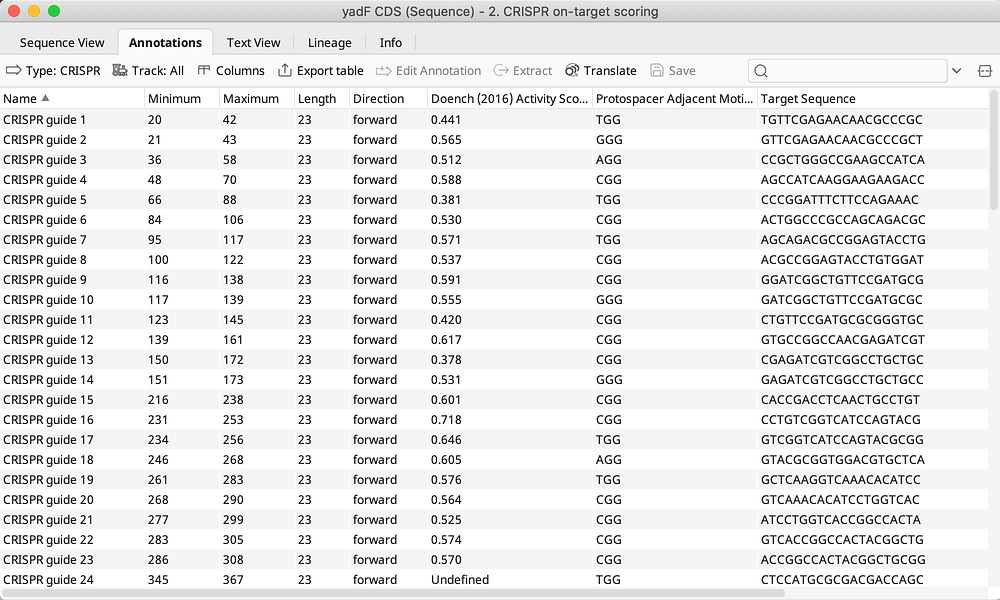
- Tabular view of candidates allows sorting and filtering of gRNAs and easy export for ordering or further analysis
- Evaluate on-target activity using the latest algorithms including Doench et al 2016
Specificity Scoring
- Confidently pick the most specific guide RNA using scoring based on Zhang et al 2013
- Check for off-interactions in regions outside the selected gene and in a database of additional sequences
- Build a custom off-target database from your own specialized or proprietary data
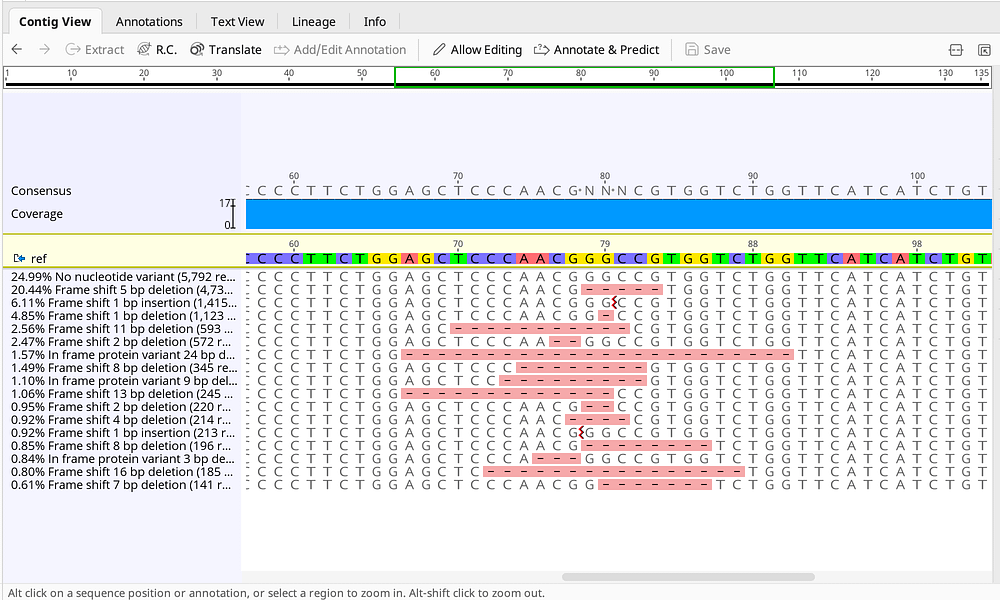
Analyze CRISPR Editing Results
- Easily align, cluster and visualize NGS reads from your CRISPR editing experiments using a purpose-built algorithm
- Analyze the frequency of variants and their protein effects from your application of choice, including HDR, NHEJ and base editors
Resources
EXPLORE GENEIOUS PRIME
Molecular Cloning
View plasmid maps, annotate vectors, find restriction sites; digest, ligate, and perform Golden Gate, Gibson, and Gateway cloning.
Primer Design
PCR and sequencing primer design tool. Primer database and testing.
Sanger Sequencing
Trim, assemble, and view Sanger sequencing trace files. Powerful SNP detection and variant calling.
Import Data
Import and convert common file types as well as their annotations and notes with a simple drag and drop
GENEIOUS ACADEMY